Tagged: translation
Protein Synthesis (part 3): Grade 9 Understanding for IGCSE Biology 3.18B
This is my final post on protein synthesis, you may be relieved to know…. It is a complicated topic and there is lots to understand but remember it is only one specification point out of several hundred in the IGCSE specification…… So don’t worry too much if you find this hard to grasp and don’t spend a disproportionate amount of time revising it. If you are fascinated by how genetic information is encoded in DNA and how genes work at a molecular level, then you have to choose Biology as a subject to study at A level!
I want to end with two final questions, both of which are essential to address if you are to acquire the grade 9 understanding you need.
What is meant by complementary base pairing and why is it so important?
Let’s go back to the structure of a DNA molecule.
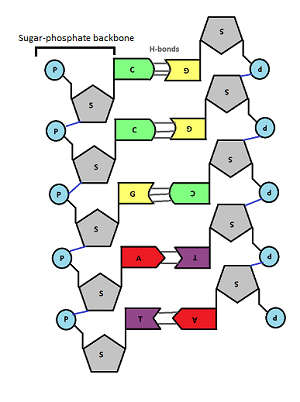
In the middle of the molecule there are pairs of bases. There are four possible bases in DNA – Adenine, Thymine, Cytosine and Guanine – but they are often represented by their letters A,T,C and G.
Bases pair up in a totally predictable way across the double stranded DNA molecule:
A always pairs with T, C always pairs with G. Why is this? Well you can see from the diagram above that A and T are held together by two weak bonds called Hydrogen (H) bonds, whereas C and G are held together by 3 H bonds. This means that this is the only way they can pair up in a stable way.
These pairs of bases (A=T and C≡G) are called complementary base pairs because they always match up in a predictable way.
You can also get complementary base pairing between RNA bases. (Remember that in RNA, there is never any thymine but it is replaced with a different base called uracil) So A=U and C≡G are the complementary base pairs between two RNA molecules.
Complementary base pairing between DNA and RNA bases is essential in transcription but you do not need to know the details at this stage.
We will come back to complementary base pairing in a moment…..
How does the ribosome ensure that the correct amino acids are joined together to make the protein during translation?
To understand this, you need to understand the role that transfer RNA (or tRNA for short) plays in protein synthesis. tRNA molecules are found in the cytoplasm and have an interesting structure. At one end, they have an important triplet of bases called the anticodon. At the other end, there is a place in the molecule where an amino acid can be added.

The key idea is this: the tRNA molecules are loaded up by attaching an amino acid by a group of enzymes found in the cytoplasm. But these enzymes ensure that each amino acid is only attached to tRNA molecules with the correct anticodon.
Transfer RNA molecules are so called because they are going to transfer (or carry) the amino acids into the ribosome. You do not need to know the details of protein synthesis but the main idea is that there is complementary base pairing between the anticodon on the tRNA and the codon on the mRNA.
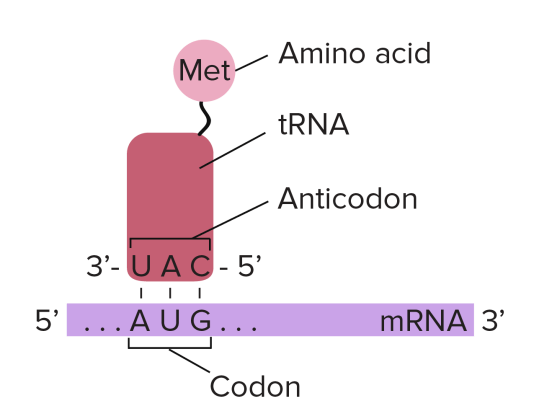
This complementary base pairing between codon and anticodon when combined with the structure of the ribosome means that the amino acids will be joined together in the correct order to make the protein.
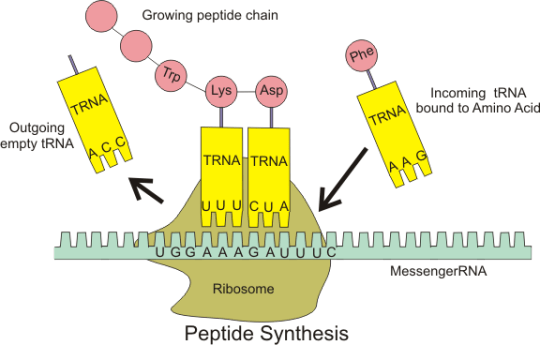
Please do not worry too much about the details of all this. I am sure that the examiners will not expect you to know the details of protein synthesis. But you should understand what an an anticodon is and the role of tRNA in the overall process.
anticodon: “a triplet of adjacent bases found on a transfer RNA molecule”
tRNA: “these molecules are found in the cytoplasm of cells and carry the correct amino acid into the ribosome. There is complementary base pairing between the codon on the mRNA and the anticodon on the tRNA as shown in the diagram above”
Protein Synthesis (part 2): Grade 9 Understanding for IGCSE Biology 3.18B
In the last post, I asked you to think of a good “2 mark” explanation of some important terms to do with protein synthesis. Here is my (GCSE level) answer…..
Gene: “a section of a DNA molecule that codes for the production of a single protein”
Ribosome: “a small structure found in the cytoplasm of cells where proteins are made by joining amino acids together into a long chain”
Transcription: “the process occurring in the nucleus in which a double-stranded DNA molecule is used to make a single-stranded molecule of messenger RNA”
Messenger RNA: ” a small single stranded molecule that is made in transcription and can carry the genetic information out of the nucleus to the ribosome”
Translation: “the second stage of protein synthesis that occurs in the cytoplasm in a ribosome in which amino acids are joined together in the correct order to make the protein”
Codon: “a triplet of adjacent bases in an mRNA molecule that codes for a single amino acid”

Finally I want to answer three important questions, one in this post, two in the next….
Why are codons three bases long?
Well, the answer here is a simple bit of Maths. If you remember, there are 20 possible amino acids that can be joined together in any order and in any number to make a protein. In DNA/RNA there are just four bases. So in order to code for 20 amino acids, how many “words” do you need? Well you need at least 20…..
In DNA/RNA you only have 4 “letters” available to make these words. (The letters are the bases and the words are the codons.)
- If the words were one letter long, there are only 4 words. (43 for the mathematicians): A,T,C,G
- If the words were two letters long, there are only 16 possible words (42 for the mathematicians): AA, AT, AC, AG, CA, CT, CC, CG, GA, GT, GC, GG, TA, TT, TC, TG
- If the words were three letters long, there are 64 possible words (43 for the mathematicians) AAA, AAT, AAC, AAG, ACA, ACT, ACC, ACG etc. etc.
So three bases in a codon is a minimum number needed to code for 20 different amino acids. But then this raises a question, doesn’t it? If there are 64 possible words and only 20 words needed, then what happens to all the extra, unnecessary codons? The answer is that there are lots of synonyms in the language of DNA/RNA. (Synonyms are two different words that have the same meaning)
Check out this picture of the genetic code (written in the language of RNA) and notice all the lovely synonyms!

Phe, Leu, Ser, Tyr, Cys and the others are names of amino acids
Notice that almost all amino acids have more than one codon that codes for them. For example, the amino acid Thr (threonine) is coded for by ACU, ACC, ACA and ACG in mRNA.
The three codons marked STOP are used as signals for the ribosome to stop the process of translation (but you don’t need to worry about that unless you wisely choose to continue your studies in Biology at A level!)
This is complicated stuff so please feel free to ask me questions about this using the “leave a comment” box below. You won’t get an instantaneous response but I try to check my blog every couple of days.
Protein Synthesis (part 1): Grade 9 Understanding for IGCSE Biology 3.18B
This is by far the most difficult concept for you to understand in the new GCSE specifications. In fact, it was only ever taught to A level students until last year (and to be honest I would much prefer it that way!). But that is no consolation to you poor folk who are going to get tested on it in your IGCSE and GCSE exams…
I am going to keep this as simple as I possibly can but am not going to dumb it down…. My blog is aimed for students who are ambitious to develop Grade 9 understanding in Biology (this topic is not tested at all in our Double Award Science course) so I want to explain it to you at the level you need. But you will need to read this carefully, take your time and you might need to break it down into small sections to build the understanding you need.
Can I suggest that first of all you read this post from my blog about DNA and how it works?
What is a gene?
A gene is a section of DNA that codes for a single protein. How does this code work? Well the short answer is that the sequence (order) of the bases as you read along the DNA molecule is a code for the sequence of amino acids that are joined together to make the protein.
Remember that there are 4 different bases in DNA: Adenine (A), Thymine (T), Cytosine (C) and Guanine (G). So a sequence of bases on a piece of DNA might look like this:
GCCTATAAATGGCAGGCATTAGCTCTAGGAAATCTAGGGACTTTACA
Protein Synthesis
Proteins are made by joining small molecules called amino acids together. This process is called protein synthesis and happens in small structures in the cytoplasm of all cells called ribosomes. But for all eukaryote cells, this poses a big geographical problem.
The “information” in the gene is stored in a sequence of bases in a DNA molecule and is found in the nucleus. DNA never leaves the nucleus because it is too important a molecule to be allowed into the reactive and unpredictable environment in the cytoplasm. But there are no ribosomes in the nucleus and these are the structures in which proteins are actually made. So a temporary intermediate molecule is needed to carry the “information” from the gene in the nucleus out into the cytoplasm where the ribosomes are found. So the process of making a protein therefore has to exist as a two stage process. The first stage is making the temporary intermediate molecule using the sequence of bases in the gene. Then there is a second stage that happens in the cytoplasm in the ribosome and this involves joining the amino acids together to make the actual protein.
This idea was called the “central dogma of molecular biology” by Watson and Crick in their famous paper on the structure of DNA.

Transcription and Translation
There is quite a lot of jargon in this topic.
Transcription is the name for the process that happens in the nucleus in which a temporary intermediate molecule is made. This temporary “information-containing” molecule is a form of RNA called messenger RNA (or mRNA for short). The mRNA travels out of the nucleus to a ribosome which is found in the cytoplasm. Here a process called Translation occurs in which the the amino acids are joined together in the correct order to make the protein.

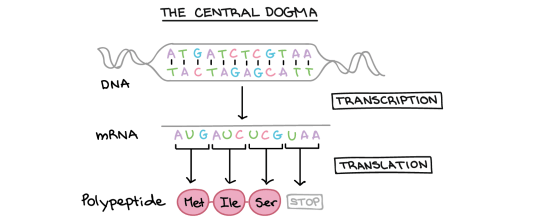
Look at this second diagram of the central dogma above. It shows a double-stranded DNA molecule at the top with pairs of bases (either A-T or C-G) joined by hydrogen bonds. The “information” in the molecule is found in the sequence of bases: on the top strand of the DNA this sequence is ATGATCTCGTAA.
You can see that transcription results in the formation of a molecule of mRNA. (Remember that RNA is always a single stranded molecule and contains the base Uracil in place of the base Thymine)
So can you see that the sequence of bases in the mRNA is almost identical to the DNA strand above, but with the base T replaced by the base U.
mRNA sequence: AUGAUCUCGUAA
This diagram shows us one final thing about how protein synthesis works. Look now at the small section of protein (polypeptide) that is produced in translation. You can see that this section of protein is made of three amino acids joined together: methionine (Met), attached to isoleucine (Ile) attached to serine (Ser)
Each amino acid is coded for by a group of 3 adjacent bases on the mRNA molecule. These triplets of bases are called Codons.
- AUG is a codon that codes for the amino acid Methionine
- AUC is a codon that codes for the amino acid Isoleucine
- UCG is a codon that codes for the amino acid Serine
(UAA is called a stop codon as it ends the translation process at the ribosome)
A codon is a triplet of adjacent bases on a mRNA molecule. Each codon codes for a single amino acid that will be joined together to make the protein.
Check your understanding:
Can you explain the meaning of the following terms? Write a 2 mark explanation of what each word means.
- Gene
- Ribosome
- Transcription
- Messenger RNA
- Translation
- Codon
I will put the answers into the next post called Protein Synthesis (part 2) which I promise I will write tomorrow…… That’s enough for now.